Circular ribonucleic acids (circRNAs) are a promising platform for gene expression studies as a stable and prevalent ribonucleic acid in eukaryotic cells, which arise from back-splicing. In a new report now published in Nature Biotechnology, Robert Chen and a team of interdisciplinary researchers at Stanford University, California, U.S., developed a systematic approach to rapidly assemble and test features affecting protein production based on synthetic circular RNAs. The team maximized translation of the circRNA by optimizing fine elements to implement design principles to improve circular RNA yield by several hundred-fold. The outcomes facilitated an increased translation of the RNA of interest, when compared to messenger RNA (mRNA) levels, to provide durable translation in vivo.
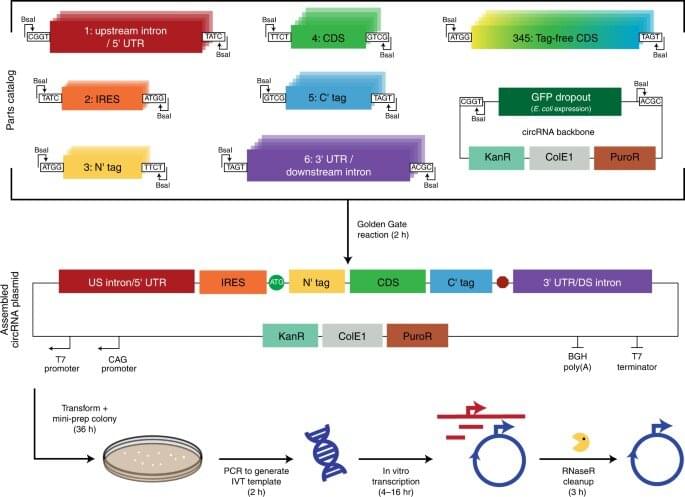
Developing circular RNA (circRNA) in the lab
Therapeutics based on ribonucleic acids span across messenger RNA (mRNA), small interfering RNAs (siRNA) and microRNAs (miRNA) with expansion into modern medicine including small molecules, biologics and cell therapeutics. For example, the lately popular mRNA vaccines can be designed in the lab and developed at a rapid pace to respond to evolving and urgent medical crises. Coding RNAs can be circularized into circRNAs to extend the duration of protein translation, based on RNA molecules that covalently join head-to-tail. Bioengineers have also advanced the synthesis of circular long transcripts into circRNAs. However, the fundamental mechanisms of initiating translation to form circular RNA or messenger RNA differ due to the lack of a 7-methylguanylate (M7G) cap on the circular RNAs. As a result of this, researchers need to thoroughly examine the principles of circular RNA translation to build better therapies and potentially surpass the translational capacities of mRNA.