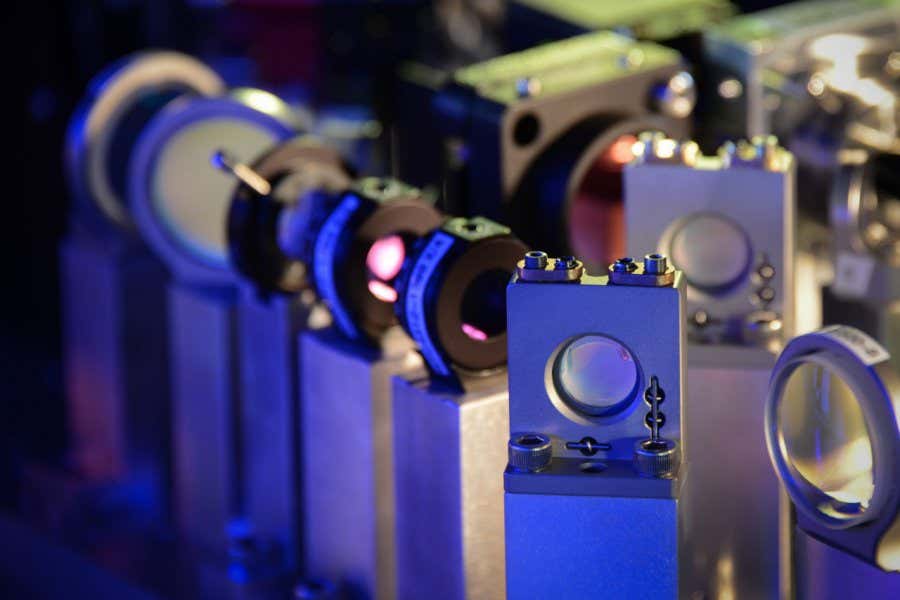
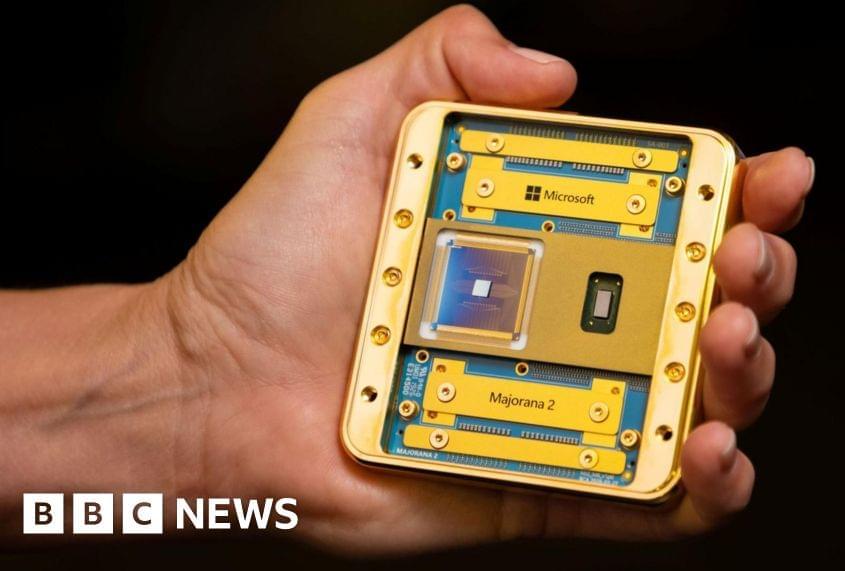
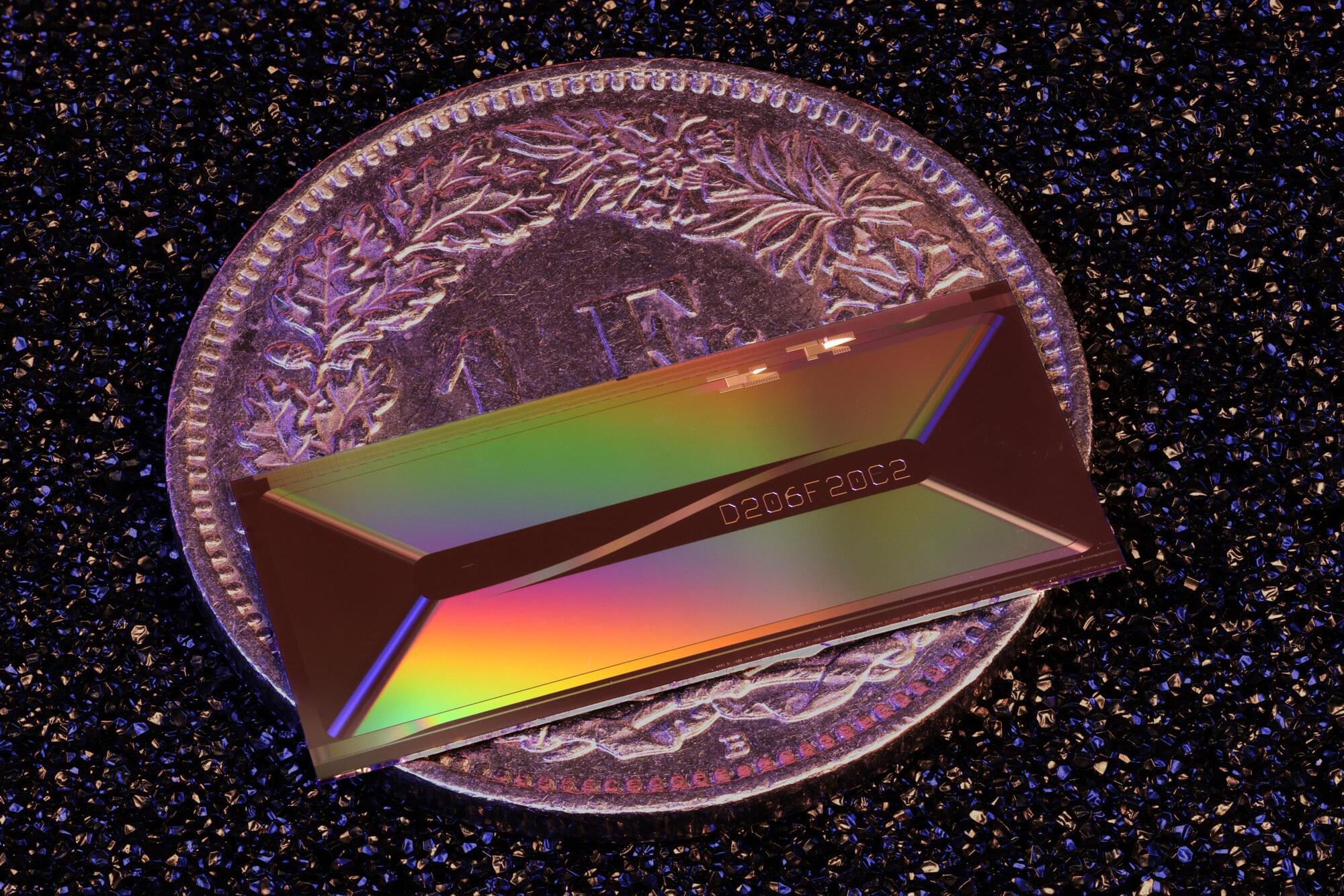
As semiconductor chips become increasingly thinner, the components inside chips are locked in a fierce race to achieve the ultimate ultra-thin state. However, this has presented a structural limitation: the thinner the device, the harder it is for electricity to flow.
Recently, a research team at POSTECH (Pohang University of Science and Technology) successfully resolved this issue through a simple yet innovative approach: “thickening only the necessary parts.”
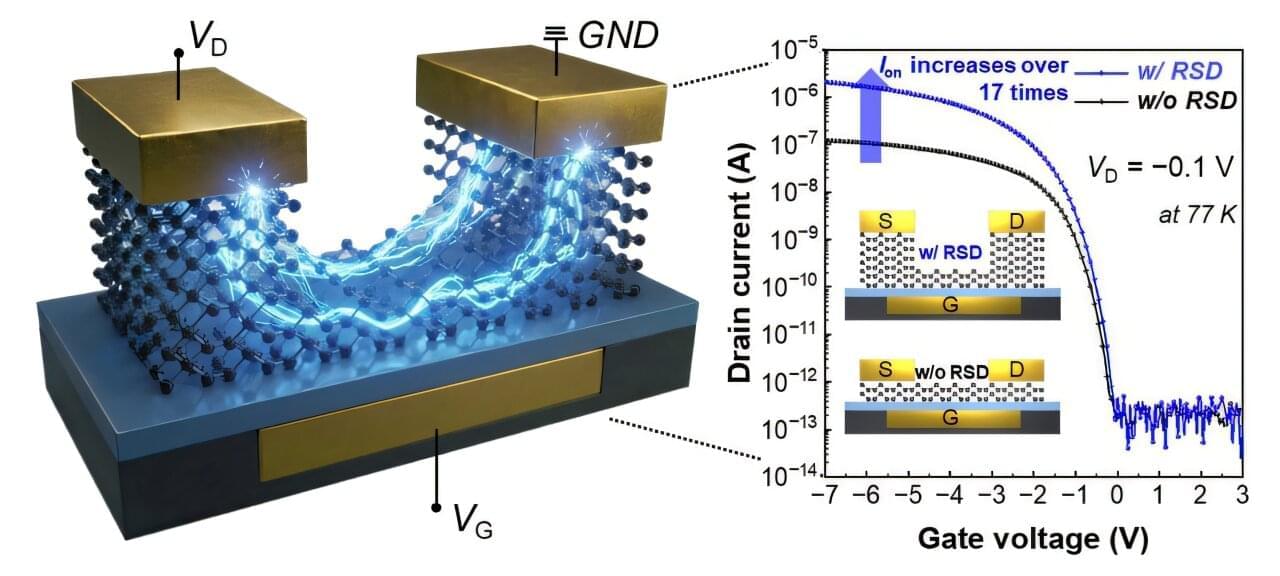
The research team, led by Professor Byoung Hun Lee from POSTECH’s Department of Electrical Engineering and the Department of Semiconductor Engineering, has developed a technology that dramatically lowers contact resistance by redesigning the metal-semiconductor contact structure in ultra-thin tellurium (Te) transistors.