Less than two decades after smartphones fit into the palm of our hands, artificial intelligence is now running on devices worn on our wrists. The challenge is that while devices continue to shrink, the amount of data they must process and the number of functions they must perform are growing exponentially. A research team at POSTECH (Pohang University of Science and Technology) has found a promising way to address this contradiction.
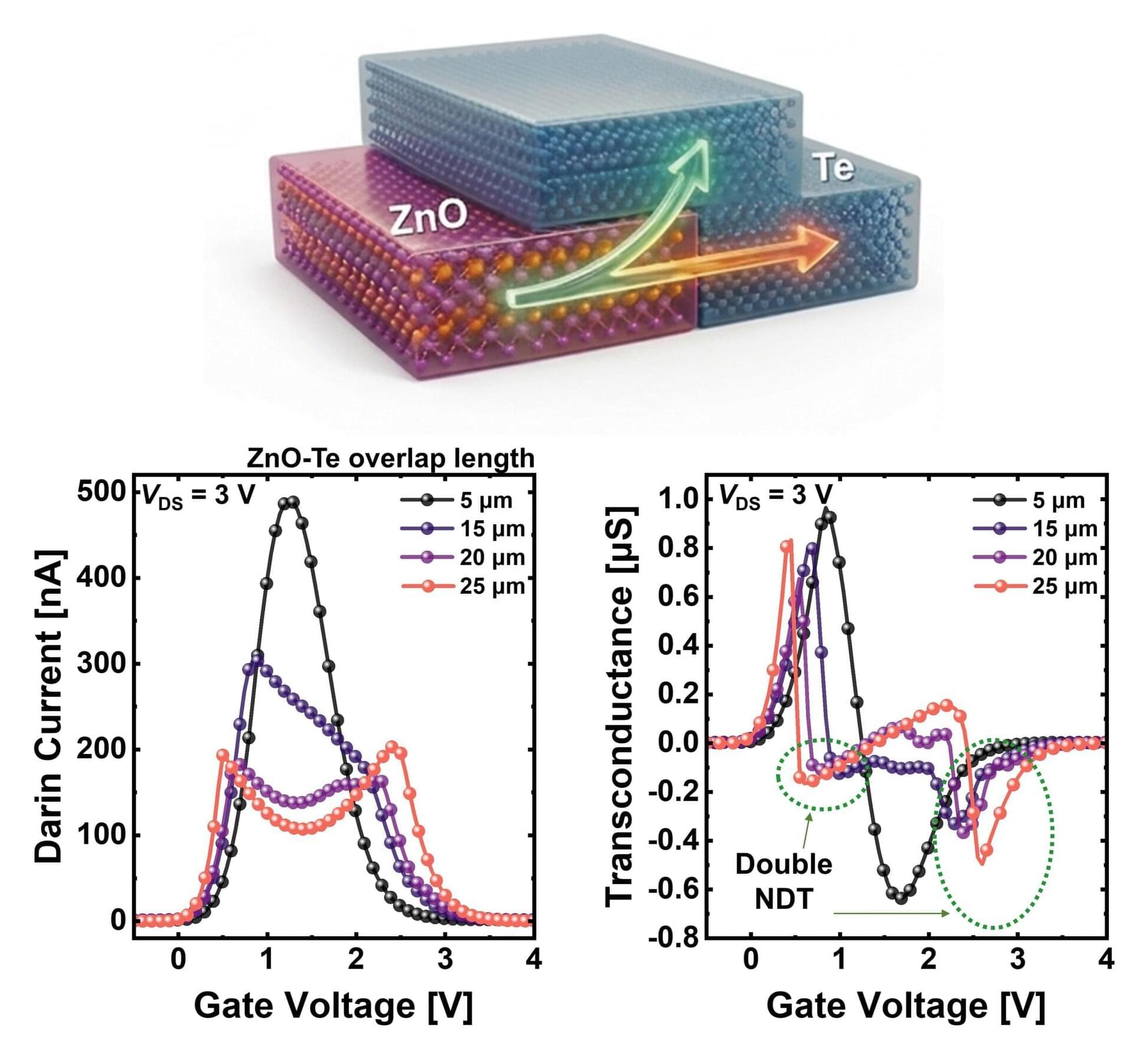
A team led by Professor Byoung Hun Lee of the Department of Electrical Engineering and the Department of Semiconductor Engineering at POSTECH, together with Dr. Jae Hyeon Jun of the Department of Electrical Engineering, has developed a transistor technology that enables a single semiconductor device to perform multiple circuit functions simultaneously. The new approach significantly simplifies circuit design and increases data processing speed fourfold compared with conventional methods. The findings were published in Advanced Functional Materials.
One of the key challenges in the semiconductor industry is integrating more functions into smaller chips. As the number of functions increases, so do the number of circuits and transistors required. However, when adding new functions to previously fabricated semiconductor chips, back-end-of-line processing must be conducted at temperatures below 400 C to protect the existing chip structure.