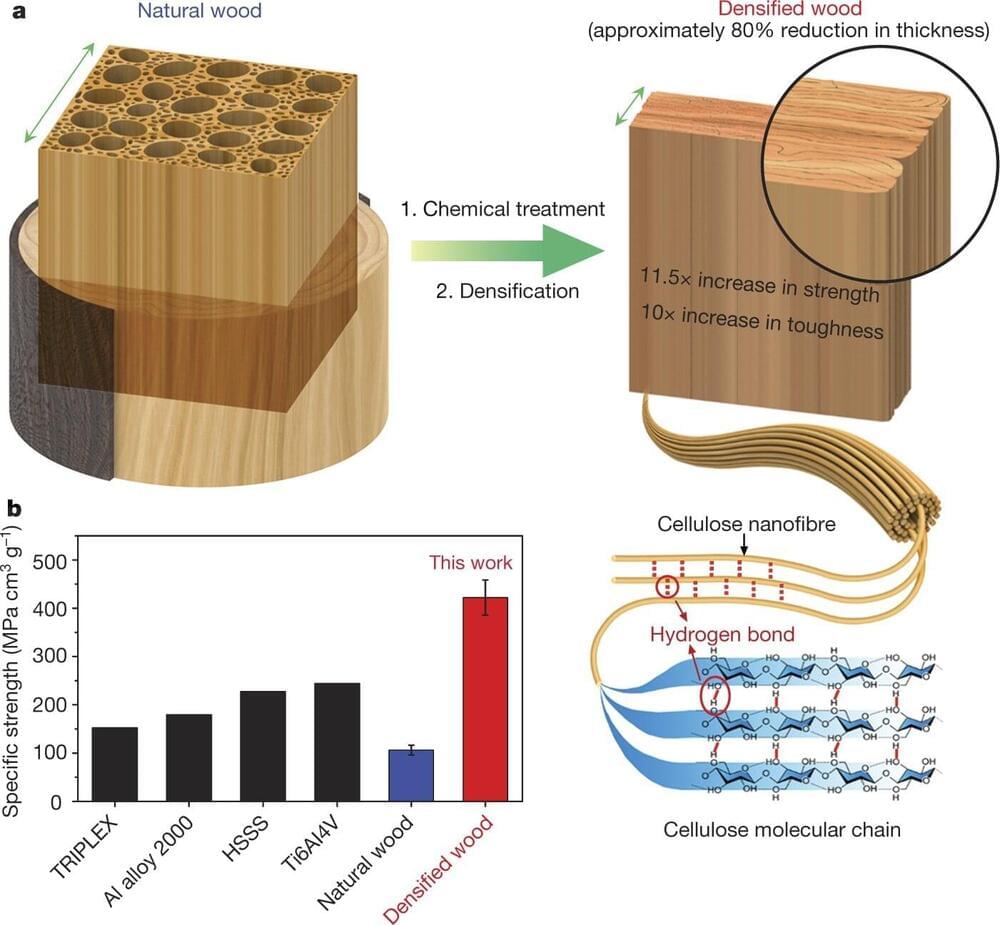
Researchers at the University of Maryland genetically modified poplar trees to produce high-performance, structural wood without the use of chemicals or energy-intensive processing. Made from traditional wood, engineered wood is often seen as a renewable replacement for traditional building materials like steel, cement, glass and plastic. It also has the potential to store carbon for a longer time than traditional wood because it can resist deterioration, making it useful in efforts to reduce carbon emissions.
But the hurdle to true sustainability in engineered wood is that it requires processing with volatile chemicals and a significant amount of energy, and produces considerable waste. The researchers edited one gene in live poplar trees, which then grew wood ready for engineering without processing.
The research was published online on August 12, 2024, in the Journal Matter.