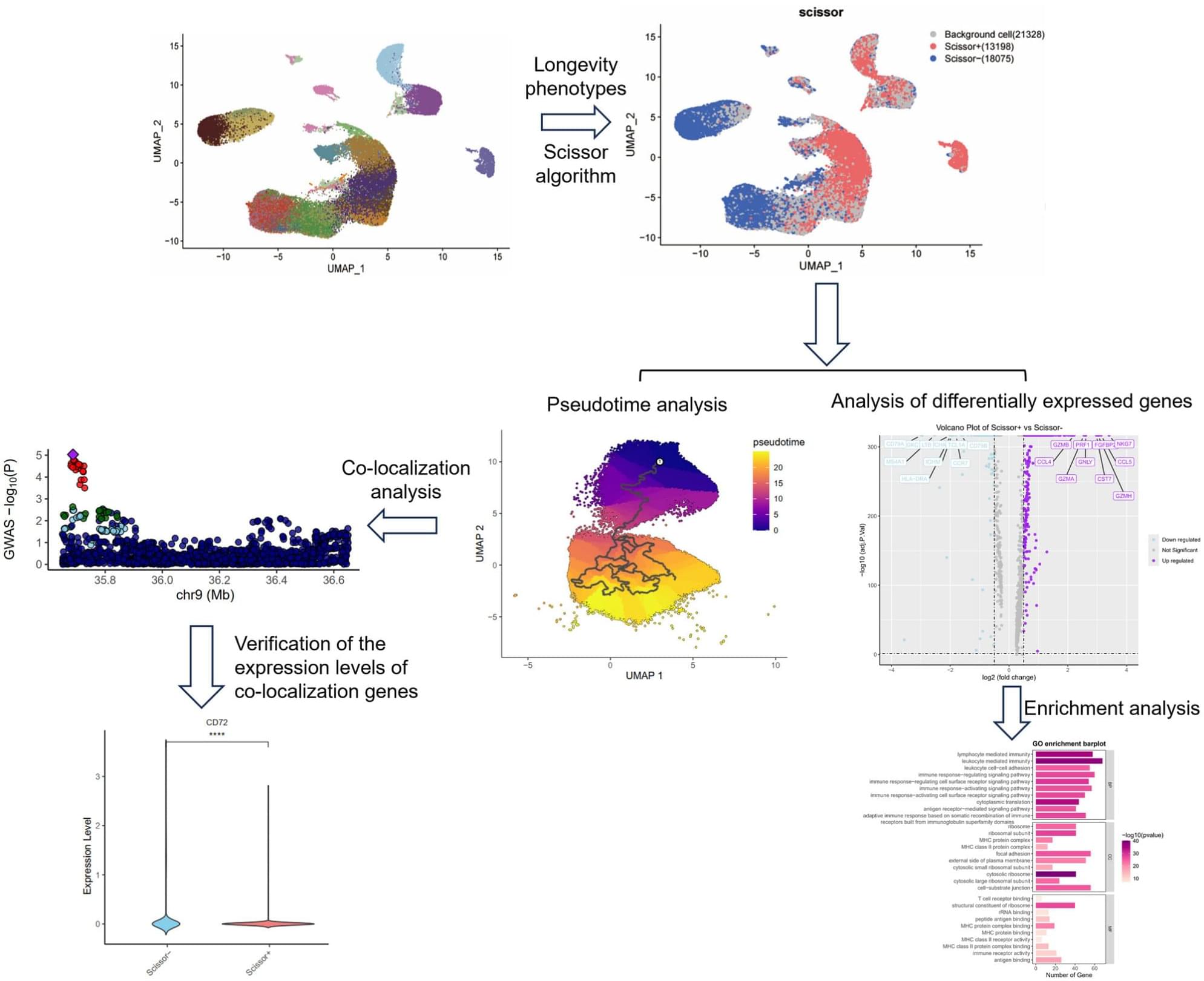
Immunosenescence represents a critical aspect of the aging process. Centenarians, serving as a nature model of “healthy aging,” demonstrate a distinctive immune “compensatory adaptation” mechanism that contributes to the maintenance of immune homeostasis. However, the specific immune cell subsets involved and the molecular mechanisms underlying these phenotypic traits remain incompletely understood. In this study, we integrated single-cell RNA sequencing data spanning the entire lifespan of East Asian populations with bulk transcriptomic data from a centenarian cohort in Guangxi. Utilizing the Scissor algorithm, we identified immune cell subpopulations positively (Scissor+) and negatively (Scissor−) associated with longevity phenotypes, thereby constructing an immune cell atlas of “Longevity Molecular Tag.” Our findings indicate that Scissor+ cells predominantly comprise natural killer (NK) cells, CD8+ T cells, and γδ T cells, characterized by enhanced cytotoxic and immunomodulatory functions. Conversely, Scissor− cells mainly include CD4+ T cells, B cells, and dendritic cells (DCs), which are linked to inflammatory signaling pathways and Th17/Th1 differentiation. Trajectory analysis elucidated the differentiation pathways of NK, CD8+ T cells, CD4+ T cells, and B cells. Differentially expressed genes were enriched in pathways such as NF-κB signaling, T cell receptor signaling, and NK cell cytotoxicity. Furthermore, co-localization analysis revealed five eQTL-colocalized events (rs3793537–GLIPR2/CD72/TLN1 and rs8019902–TRDV2/TRDC) associated with longevity. Collectively, these results suggest that centenarians achieve immune equilibrium by remodeling cytotoxic immune lineages and finely tuning inflammatory responses, thereby promoting health span and longevity. This study offers novel insights into potential strategies for modulating immunosenescence.